Can a General LLM Diagnose a DICOM Slice?
The answer is absolutely fucking not. The industry is optimizing the wrong layer. A general multimodal LLM is not replacing a radiologist any time soon, and even a domain-trained model is not a substitute for clinical responsibility. The useful role is narrower: review, evidence gathering, drafting, and workflow support. The real question is operational cost: large DICOM studies, expensive inference, strict reliability, and regulated data handling.
This is a real benchmark, but it is still small. It is not a clinical validation study and it does not support diagnostic claims. It answers one narrow question: how far can a general multimodal model go on a public single-slice DICOM task before confidence stops matching reality?
What I Tested
I used 10 public Pacsbin teaching cases: 7 CT or CTA studies and 3 MRI studies. For each case I selected one pathology-bearing key image, downloaded the linked DICOM instance, rendered it with the case page's own window settings, and asked the model for the single most likely diagnosis visible on that slice. The score was simple. A response was a match, a partial, or a miss.
What Happened
The result was weak. Strict top-1 accuracy was 3/10. Two more answers were directionally useful but incomplete, which brought the softer score to 5/10. The misses were not random. The model often locked onto a structure that was visually obvious but clinically secondary, then built a confident story around the wrong disease. The ugliest example was acute appendicitis misread as bilateral osteitis condensans ilii at 0.89 confidence. It sounded convincing. It was obviously wrong.
What Changed In The Follow-up
The original public benchmark used the Codex CLI path on gpt-5.4. The follow-up harness also
included direct OpenAI API testing, and the published review-pipeline comparison used
gpt-4.1-mini. The new workflow was ROI-first and stopped trusting one answer. One pass made the
first call, another proposed a competing explanation, and a few checks tried to decide which story held up
better. That made the output easier to inspect and audit. It still did not solve the core problem.
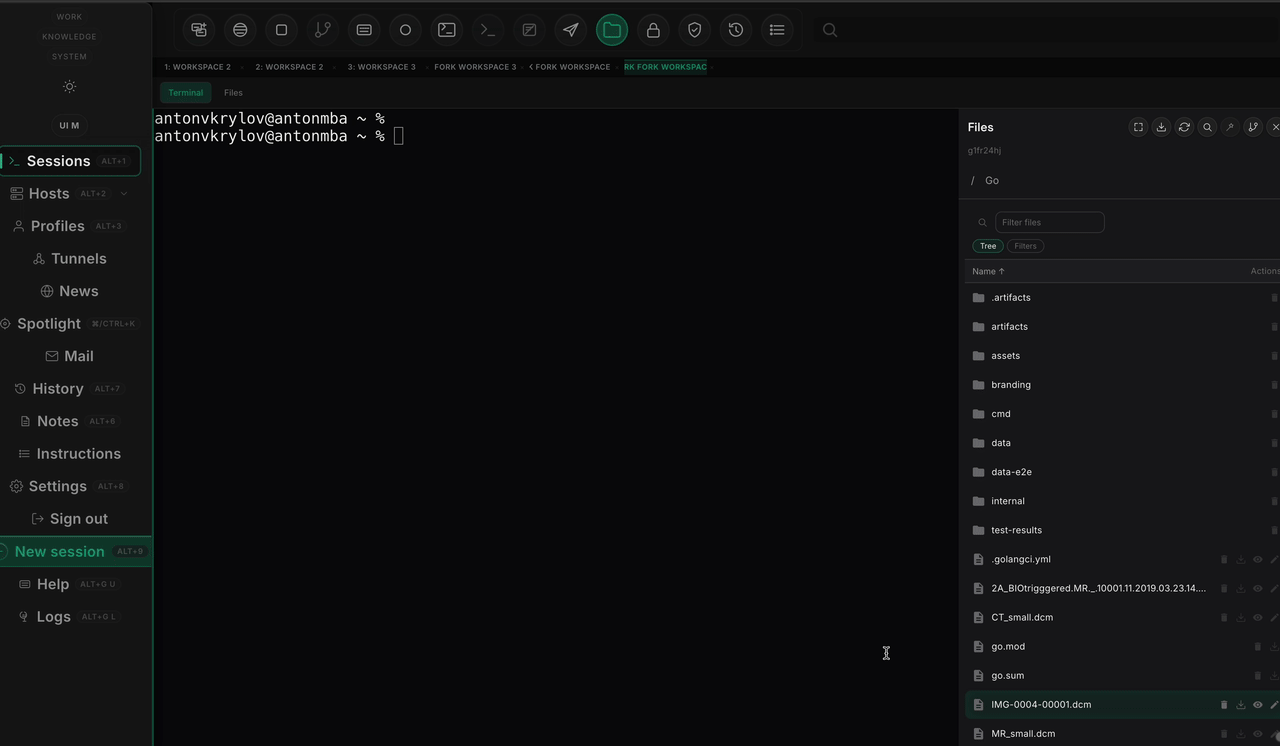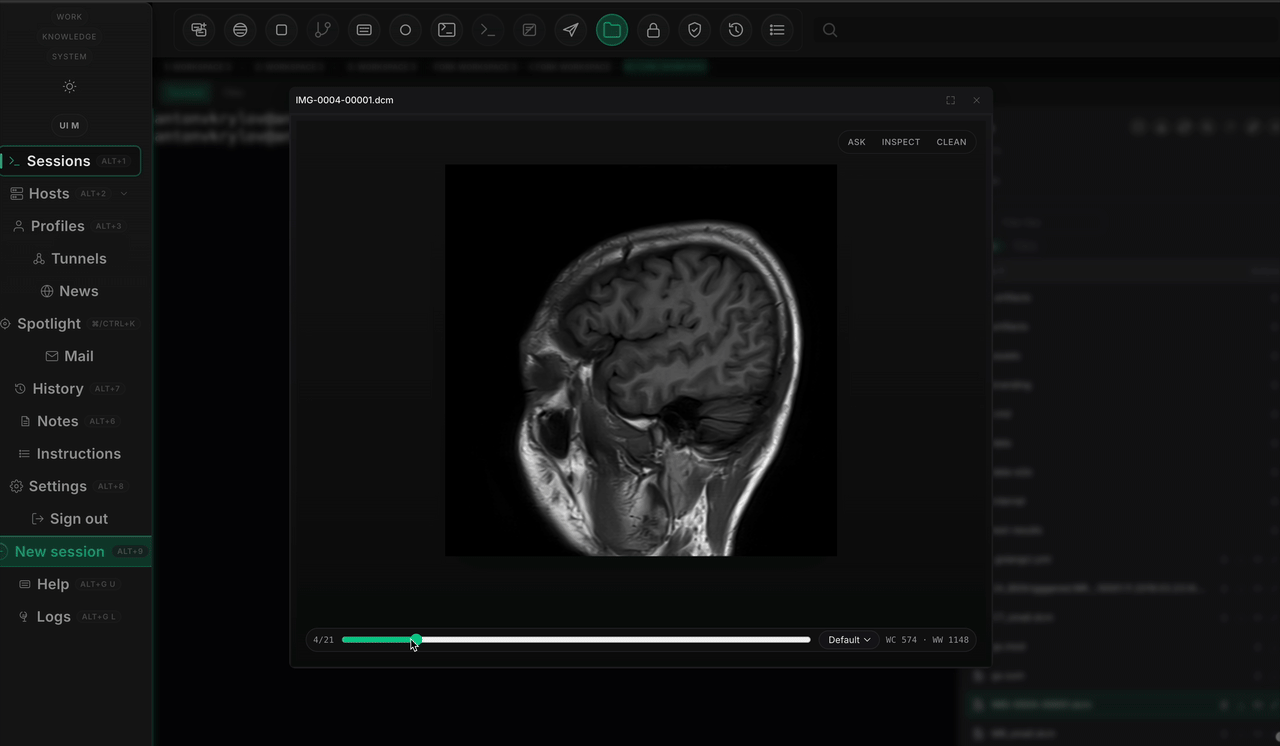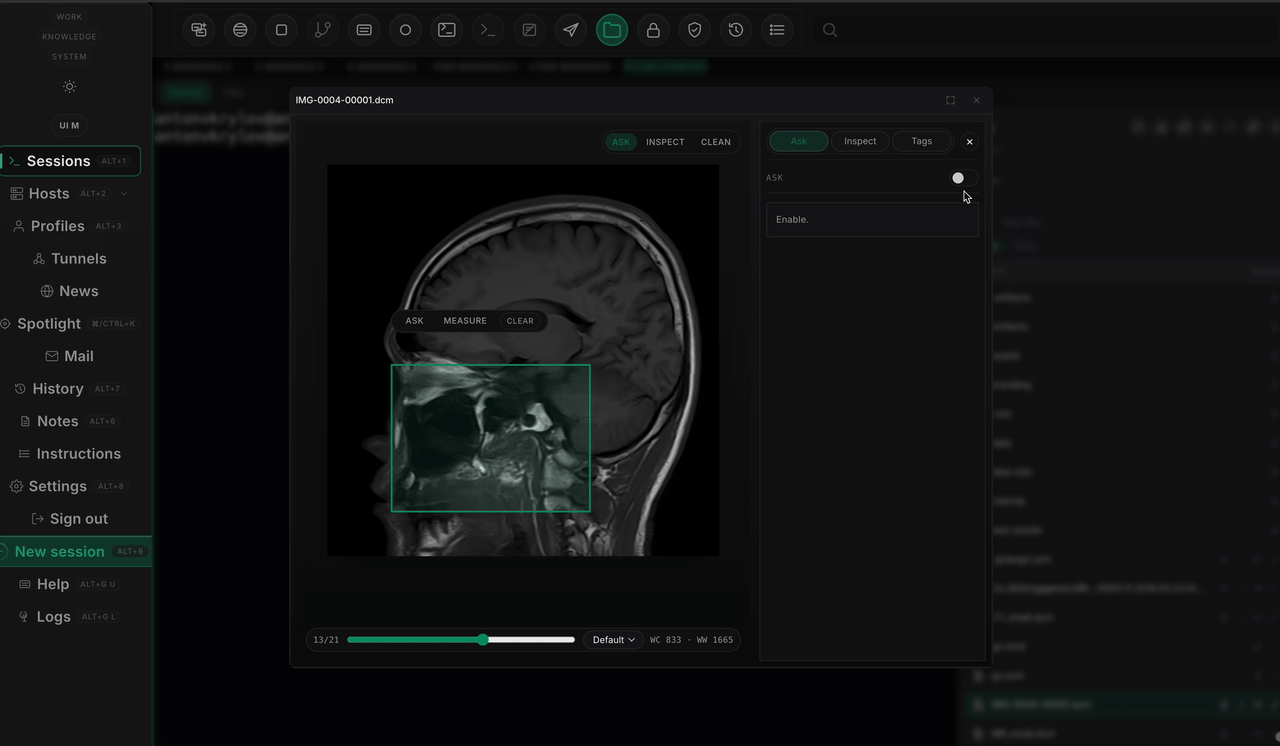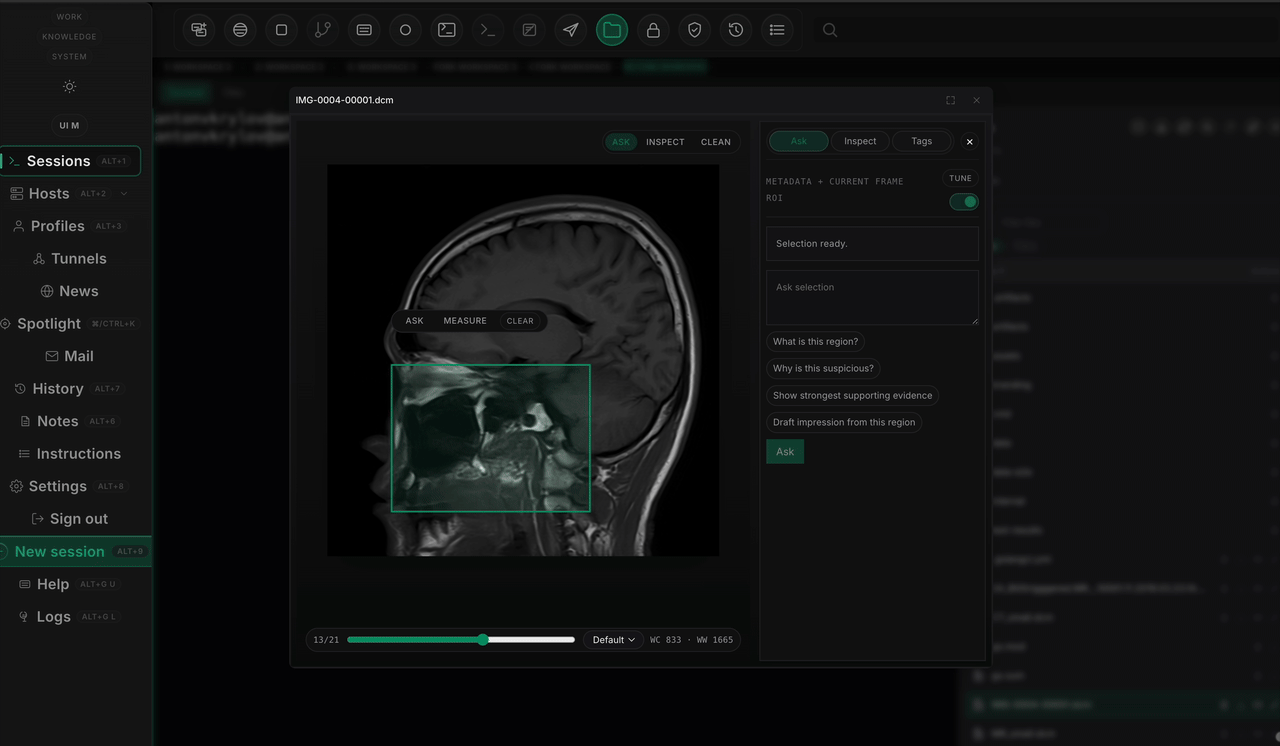
The second benchmark made the lesson sharper. ROI guidance helped. Extra review logic helped the system explain itself. But the review pipeline still did not beat the same-run finder baseline on the fairest comparison. Reasoning improved faster than perception.
Operational Cost
Medical AI becomes an operational problem very quickly. A DICOM study is not a neat image file. It can span hundreds or thousands of instances and reach gigabytes. In many regulated industries, public cloud is not a free choice, so pushing that data through laptops, temp folders, and shared drives is slow, expensive, and hard to control.
A concrete example is a worker pod reading a large study from mounted storage. If the network share stalls or drops, the worker can fail in the middle of processing. In a script-like setup, that often leaves the case hanging in an unclear state. In a fault-tolerant clustered runtime, the pod can die, restart, or be rescheduled while the case state stays outside the process in a queue or status store. The same applies to scale: metadata workers and heavy GPU inference workers can scale separately instead of fighting for one machine. That is the real value. Not that failures disappear, but that storage faults, restarts, and load spikes stop breaking the whole pipeline.
This matters even more because medical environments are highly regulated, often tied to legacy PACS and hospital software, and usually built on a fragmented stack of old viewers, file shares, vendor gateways, and custom interfaces. That is why localhost is the wrong center of gravity. The safest path is to keep files, models, and workers inside the controlled runtime and use the browser only as the control surface.
For users, that changes daily work. They stop moving data through their laptop just to inspect or patch something inside a pod. The browser becomes the control surface and the runtime stays where the files and workers already live. That means fewer broken local environments, fewer kube-context mistakes, fewer copied secrets, and less context switching between terminal, editor, and file browser. It also makes failure handling much safer: atomic saves, snapshots, rollbacks, live watch streams, and controlled transfers replace a lot of “scp and hope.” A related version of the same argument shows up in the previous post on Portal Long-Term Memory.
Conclusion
The conclusion is simple. The real bottleneck is not the radiologist, and replacing radiologists with LLMs is not a credible near-term goal. The model is not the product. The operational surface is. In this benchmark the model was sometimes useful, often articulate, and still wrong too often to earn diagnostic trust. The realistic opportunity is replacing the brittle legacy software stack around the reader with better systems for review, evidence, drafting, and workflow.
If you want to inspect the work, the evaluation script, the raw benchmark JSON, and the review-pipeline comparison JSON are all public.